New TRAPP webserver for studying TRAnsient Pockets in Proteins
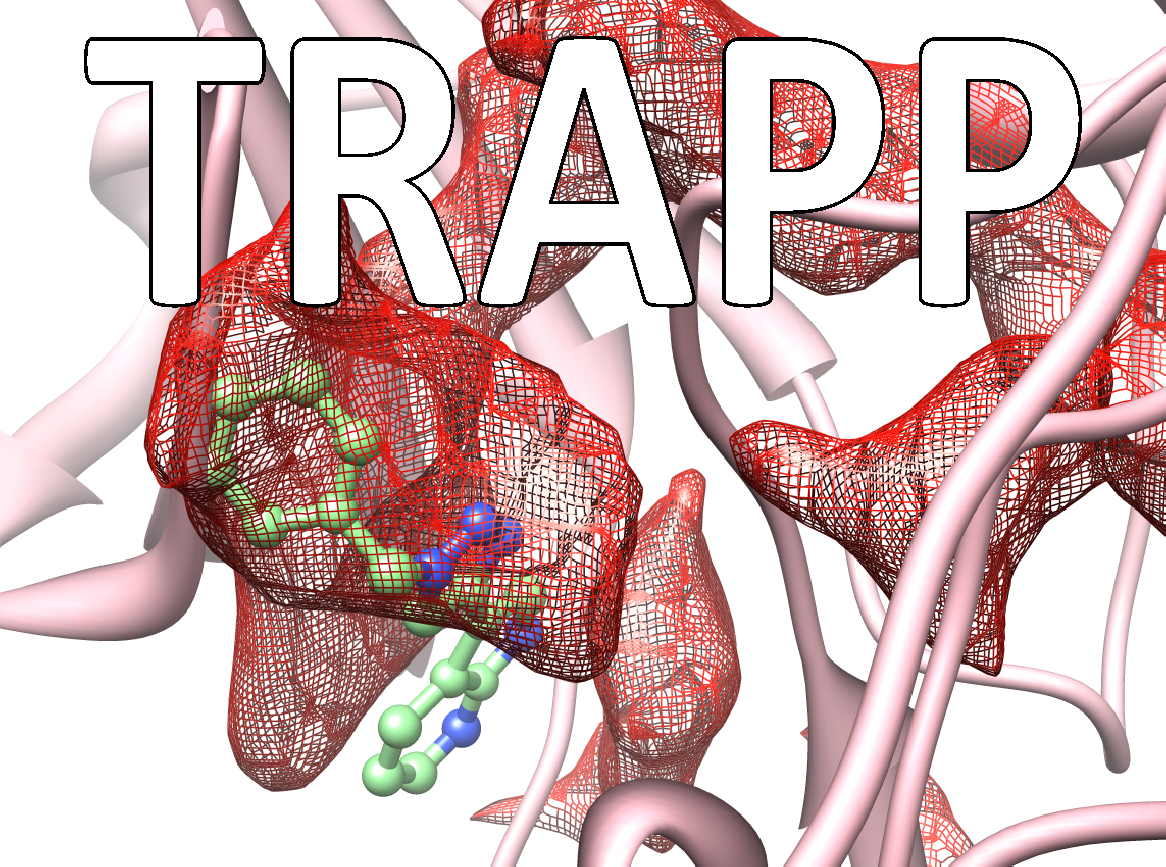
Many drugs exert their therapeutic effects by binding in cavities or pockets on specific target protein macromolecules. The binding pockets on proteins are mobile and dynamic and constantly changing shape. Some pockets or subpockets can be described as transient because they are not always present in the given protein. In structure-based drug design procedures, the dynamic nature of the target protein is usually neglected. A crystal structure of the protein may not reveal transient pockets as it provides a picture of a fixed state of the protein. The analysis of molecular dynamics simulations of a protein can however reveal them. Transient pockets are particularly interesting because they can open and close, allowing ligands to bind. Transient pockets can therefore be exploited to design compounds with greater target specificity or better kinetic binding properties than would be achieved by considering the target as a static structure.
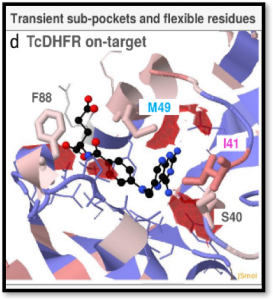
The TRAnsient Pockets in Proteins (TRAPP) webserver, developed in the Molecular and Cellular Modeling (MCM) group at HITS and recently published in the journal Nucleic Acids Research, allows users to employ a range of computational methods to generate snapshots of protein structures, to evaluate them for flexibility, and to detect and characterize transient pockets. The information TRAPP provides can be used to explore protein motion and flexibility, the appearance of transient pockets and the physiochemical, sequence and functional characteristics of binding pockets. The TRAPP webserver provides a user-friendly environment for a dynamic-structure-based approach to drug discovery.
Publication:
Antonia Stank, Daria B. Kokh, Max Horn, Elena Sizikova, Rebecca Neil, Joanna Panecka, Stefan Richter and Rebecca C. Wade. TRAPP webserver: predicting protein binding site flexibility and detecting transient binding pockets. Nucleic Acids Res (2017) gkx277. (Fulltext)
About HITS
HITS, the Heidelberg Institute for Theoretical Studies, was established in 2010 by physicist and SAP co-founder Klaus Tschira (1940-2015) and the Klaus Tschira Foundation as a private, non-profit research institute. HITS conducts basic research in the natural, mathematical, and computer sciences. Major research directions include complex simulations across scales, making sense of data, and enabling science via computational research. Application areas range from molecular biology to astrophysics. An essential characteristic of the Institute is interdisciplinarity, implemented in numerous cross-group and cross-disciplinary projects. The base funding of HITS is provided by the Klaus Tschira Foundation.