The best of both worlds
KIMMDY is born! A new simulation scheme combining Molecular Dynamics (MD) simulations with Kinetic Monte Carlo (KMC) enables bond scissions in all-atom simulations. The advantage of this hybrid approach over previous reactive simulation methods lies especially in its computational efficiency. Several orders of magnitude in timescales are bridged between the input sampling in MD and the reaction in the KMC step. This method developed at HITS has now been published in „Journal of Chemical Theory and Computation“.
Proteins in their natural environment are exposed to various mechanical loads that might lead to covalent bond scissions even before macroscopic failure occurs. In regular Molecular Dynamics (MD) simulations, covalent bonds are predefined and chemical reactions cannot occur. Furthermore, such events rarely take place on timescales accessible with MD. Existing reactive approaches either rely on computationally expensive quantum calculations (e.g. QM/MM) or complex bond order formalisms in force fields (e.g. ReaxFF).
The new simulation scheme called KIMMDY (Kinetic Monte Carlo / Molecular Dynamics), developed from the Molecular Biomechanics group at HITS, can alleviate these issues through a hybrid combination of different simulation methods: bond rupture rates are calculated based on the interatomic distances in the MD simulation and then serve as an input for a Kinetic Monte Carlo step. This approach bridges the often apparent separation of accessible timescales.
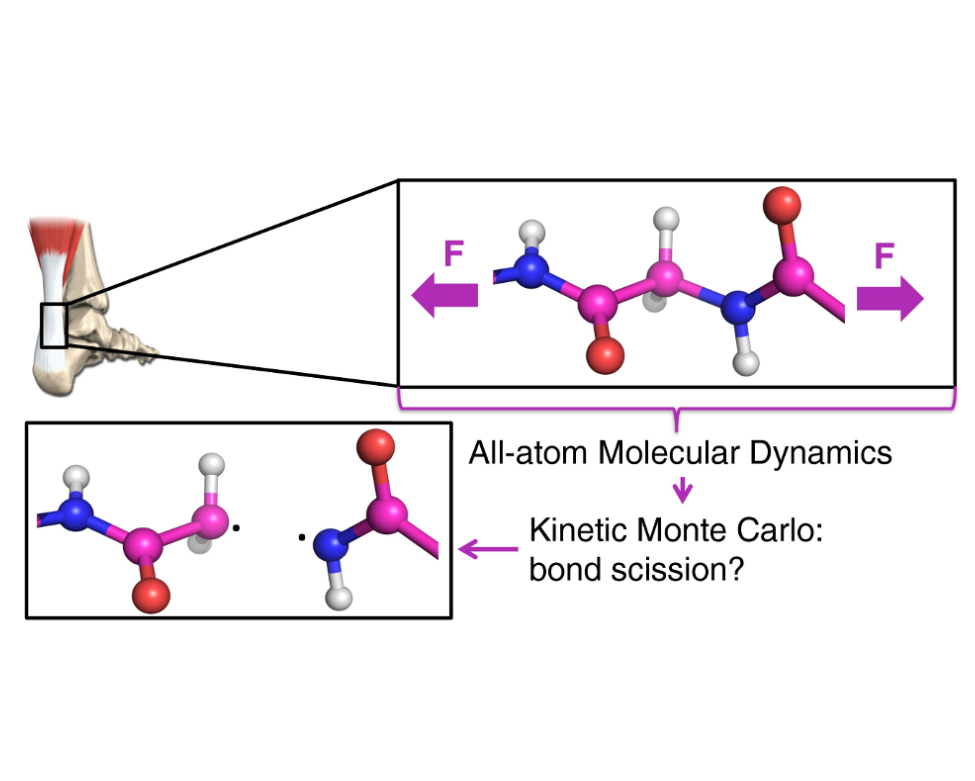
With this new technique bond ruptures in multi-million atom systems can now be investigated. Using KIMMDY it was shown that in tensed collagen, a structural protein of skin, bones and tendons, bond scissions clearly concentrate near the chemical crosslinks. After a (homolytic) bond breakage due to high tensile stress, highly reactive radicals are created in collagen. The subsequent created species could potentially act in signaling processes, converting mechanical forces into oxidative stress. Hence, this method opens up the path to further examine this biological highly relevant mechanism. It is also straightforwardly applicable to other complex (bio)materials under load and related chemistries.
The paper originated from the master thesis of scientist Benedikt Rennekamp at HITS and Heidelberg University. Since August 2019 he is PhD Student at HITS in the group of Frauke Gräter and a member of the Max Planck School «Matter to Life».
The most recent version of the software code is available online here.
Article in Journal of Chemical Theory and Computation:
Hybrid Kinetic Monte Carlo / Molecular Dynamics Simulations of Bond Scissions in Proteins
Benedikt Rennekamp, Fabian Kutzki, Agnieszka Obarska-Kosinska, Christopher Zapp, and Frauke Gräter
Journal of Chemical Theory and Computation
DOI: 10.1021/acs.jctc.9b00786
Scientific Contact:
Prof. Dr. Frauke Gräter
Group Leader „Molecular Biomechanics“
HITS – Heidelberg Institute for Theoretical Studies
E-mail: frauke.graeter@h-its.org
About HITS
HITS, the Heidelberg Institute for Theoretical Studies, was established in 2010 by physicist and SAP co-founder Klaus Tschira (1940-2015) and the Klaus Tschira Foundation as a private, non-profit research institute. HITS conducts basic research in the natural, mathematical, and computer sciences. Major research directions include complex simulations across scales, making sense of data, and enabling science via computational research. Application areas range from molecular biology to astrophysics. An essential characteristic of the Institute is interdisciplinarity, implemented in numerous cross-group and cross-disciplinary projects. The base funding of HITS is provided by the Klaus Tschira Foundation.